x <- Sys.Date()
class(x)
#> [1] "Date"24 S3, S4, and S7
You call print(x) and it does the right thing whether x is a number, a data frame, or a linear model. How? R looks at the class of x and picks the right print function. That is dispatch.
24.1 What dispatch means
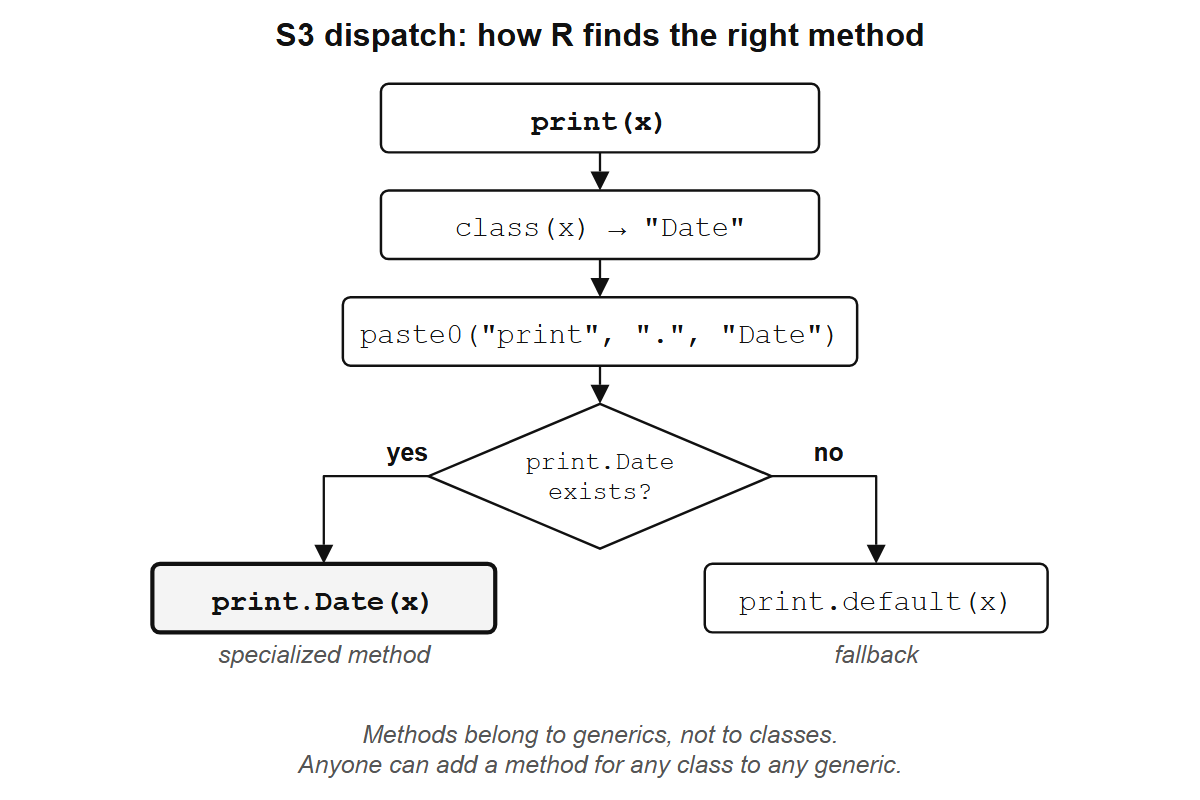
print(x) isn’t one function. It’s a generic: a function that inspects its input and delegates to a specialized method. print.data.frame, print.Date, print.lm: each is a method. The generic print finds the right one based on class(x).
When you call print(x), R sees that class(x) is "Date", looks for a function named print.Date, and calls it. If no such function exists, R falls back to print.default.
summary(), plot(), mean(), [: all generics. R is full of them. You can list the methods for any generic:
head(methods(print), 10)
#> [1] "print.acf" "print.activeConcordance"
#> [3] "print.AES" "print.anova"
#> [5] "print.aov" "print.aovlist"
#> [7] "print.ar" "print.Arima"
#> [9] "print.arima0" "print.AsIs"This is functional OOP: methods belong to generics, not to classes. You call print(x), not x.print(). In Java, Python, or C++, methods belong to objects and you write object.method(). R dispatches on the first argument. Anyone can add a method for any class to any generic without modifying the class. Package A defines a class, package B defines a method for it, and neither needs to know the other exists.
Dispatch is R picking which function to call, the same principle as “everything that happens is a function call” (Section 7.5).
In type theory, the type of an expression determines which reduction rules apply. S3 dispatch is the same idea: the class attribute determines which method runs, and the generic is a polymorphic term whose behavior is resolved at runtime by inspecting the type of its argument. This is ad-hoc polymorphism (different code for different types), as opposed to the parametric polymorphism of Hindley-Milner type systems, where the same code works for all types. R does not have parametric polymorphism in the formal sense, but functions like length() and c() behave parametrically: they work on any vector regardless of element type.
S3 dispatch also connects to sum types. When you give an object a class, you are tagging it as one variant of a type. class(x) = "penguin" says “this object is a penguin, not a dog, not a book.” The generic function then pattern matches on the tag, dispatching to the method for that variant, exactly how sum types work in Haskell (case x of) or Rust (match x): inspect the tag, run the corresponding branch. R’s mechanism is informal (a string attribute instead of a compiler-checked type), but the algebraic structure is the same sum type you saw with TRUE | FALSE (Section 8.1) and factor levels (Section 12.4). The class vector is the tag; UseMethod() is the match.
When you call print(x) and R inspects class(x) to choose a method, the class attribute is playing the same role that a type annotation plays in a statically typed language: it determines which code runs. In Haskell, the compiler verifies at compile time that every case is handled. In R, you find out at runtime that no method exists for your class, and print.default picks up the slack. That informality gives you freedom — attaching a class string to any object without declaring a type hierarchy first is useful in interactive work, where data structures evolve faster than formal definitions. But the freedom costs you exhaustiveness: every missing method is a bug that only surfaces when someone passes an object you did not anticipate.
24.2 S3 basics
S3 is informal. A class is just a string attached to an object:
x <- structure(list(name = "Adelie", mass = 3750), class = "penguin")
class(x)
#> [1] "penguin"That’s all it takes: a list (or vector) with a class attribute attached to it.
A generic is a function that calls UseMethod():
greet <- function(x, ...) UseMethod("greet")A method is a function named generic.class:
greet.penguin <- function(x, ...) paste("Hello,", x$name)
greet(x)
#> [1] "Hello, Adelie"The dispatch path: greet(x) checks class(x), finds "penguin", looks for greet.penguin, calls it. If no method is found, R falls back to greet.default (if you defined one).
You can watch this happen with sloop::s3_dispatch():
sloop::s3_dispatch(greet(x))
#> => greet.penguin
#> greet.defaultThe => arrow marks the method that was actually called. The * marks methods that exist but weren’t called. This is useful for debugging inheritance chains (more on that in Section 24.4).
Exercises
- Create an S3 class
"dog"with fieldsnameandbreed(usestructure()and a list). Write aprint.dogmethod that prints something like"Rex (Labrador)". Verify thatprint(your_dog)calls your method. - Write a
greet.defaultmethod that returns"I don't know how to greet this". Test it on a plain list without a class. - Use
sloop::s3_dispatch()to see the dispatch path forprint(Sys.Date()). Which method gets called?
24.3 Writing S3 classes properly
Setting a class with structure() works, but it’s fragile. Nothing stops someone from creating a "penguin" with a numeric name or negative mass. The established best practice is three functions: a constructor, a validator, and a helper.
The constructor (new_penguin) creates the object with minimal checks. It is for developers:
new_penguin <- function(name, mass) {
structure(list(name = name, mass = mass), class = "penguin")
}The validator (validate_penguin) checks invariants. It ensures the object makes sense:
validate_penguin <- function(x) {
if (!is.character(x$name) || length(x$name) != 1) {
stop("`name` must be a single character string")
}
if (!is.numeric(x$mass) || length(x$mass) != 1 || x$mass <= 0) {
stop("`mass` must be a single positive number")
}
x
}The helper (penguin) is user-facing. It calls the constructor, then the validator:
penguin <- function(name, mass) {
validate_penguin(new_penguin(name, mass))
}Now compare:
# Direct construction: no safety net
bad <- structure(list(name = 42, mass = -100), class = "penguin")
# Helper: catches the problem
penguin(42, 3750)
#> Error in `validate_penguin()`:
#> ! `name` must be a single character stringThe constructor is fast and trusting. The validator is thorough. The helper is what users call. This separation of concerns comes from Advanced R: construction is cheap, validation is careful, and the user interface is clean.
If you write S3 classes, write all three functions.
Exercises
- Write the constructor/validator/helper trio for a
"book"class with fieldstitle(character),pages(positive integer), andyear(four-digit number). Test the validator by trying to create a book with -50 pages. - Add a
print.bookmethod that prints something like"Title (Year, N pages)".
24.4 Inheritance
S3 supports single inheritance via class vectors:
emperor <- structure(
list(name = "Emperor", mass = 23000, dive_depth = 500),
class = c("diving_penguin", "penguin")
)
class(emperor)
#> [1] "diving_penguin" "penguin"When you call greet(emperor), R looks for greet.diving_penguin first, then greet.penguin, then greet.default. The class vector is a priority list.
sloop::s3_dispatch(greet(emperor))
#> greet.diving_penguin
#> => greet.penguin
#> greet.defaultNo greet.diving_penguin exists, so R falls through to greet.penguin. You can define a specialized method and use NextMethod() to delegate to the parent:
greet.diving_penguin <- function(x, ...) {
parent_greeting <- NextMethod()
paste(parent_greeting, "(dives to", x$dive_depth, "m)")
}
greet(emperor)
#> [1] "Hello, Emperor (dives to 500 m)"NextMethod() calls the next method in the chain, like super() in Python or Java. The diving penguin method adds dive information to whatever the penguin method already produces.
Keep inheritance shallow. One or two levels is typical in R. Deep hierarchies (five classes deep, overriding methods at every level) are rare and usually a sign you want a different design. If you find yourself building a tree of classes, step back and ask whether composition (storing one object inside another) would be simpler.
Exercises
- Create a
"fiction_book"class that inherits from"book"(from the previous exercise) and adds agenrefield. Write aprint.fiction_bookmethod that usesNextMethod()and appends the genre. - Use
sloop::s3_dispatch()onprint(your_fiction_book)to see the dispatch chain.
24.5 Why S3 works
S3 is R’s dominant OOP system, and the name tells you where it came from. When John Chambers designed the S language at Bell Labs in the early 1980s, he needed a way to make print() and summary() do different things for different kinds of objects. The solution was deliberately informal: attach a class string to any object, and let the generic function look up the right method by pasting the generic name and the class name together. No class registry, no formal type hierarchy, no compiler checks. Just a naming convention. This was S version 3, so the system became known as S3.
It was supposed to be temporary. By 1998, Chambers had designed S4, a formal OOP system with typed slots, multiple dispatch, and a class registry enforced by setClass() and setGeneric(). S4 was adopted by the Bioconductor project for its genomics infrastructure, but the rest of the R community kept using S3. S3 let package authors add methods to existing generics without coordination, which is exactly what a decentralized ecosystem of 20,000+ CRAN packages needs.
Three decades later, S7 arrived. A joint working group from R-Core, Bioconductor, and the tidyverse designed it to combine S3’s simplicity with S4’s safety: typed properties, built-in validation, clean syntax, and full backward compatibility with S3 dispatch.
S3 requires no registration, no formal definitions, and no boilerplate. You can add methods to existing generics from any package: write print.my_class in your package and it just works. You can prototype quickly: slap a class on a list and start writing methods.
The downside is obvious: no enforcement. Nothing stops you from creating a "penguin" with a numeric name or negative mass, unless you write the validator yourself. S3 trusts the programmer.
S3 works because R programmers are disciplined. Follow the constructor/validator/helper pattern and you get most of the safety of a formal system with none of the ceremony. Skip the pattern and you get bugs that hide until production.
24.6 S4 in brief
S4 adds formality. setClass() defines a class with typed slots, setGeneric() creates a generic, and setMethod() registers a method:
setClass("Animal",
slots = list(
name = "character",
mass = "numeric"
)
)
setGeneric("describe", function(x, ...) standardGeneric("describe"))
setMethod("describe", "Animal", function(x, ...) {
paste(x@name, "weighs", x@mass, "g")
})
a <- new("Animal", name = "Cat", mass = 4500)
a@name
describe(a)Slots are accessed with @ instead of $. Validation can be built in via the validity argument to setClass().
S4’s distinctive feature is multiple dispatch: setMethod("combine", c("Matrix", "vector"), ...) chooses a method based on the types of both arguments, not just the first. S3 dispatches on one argument only and cannot naturally express “when type A meets type B, do C.” Julia (2012) made multiple dispatch its central paradigm. The trade-off is real: single dispatch is simpler, multiple dispatch is more expressive but harder to reason about.
S4 is used extensively in Bioconductor and some base R infrastructure (the Matrix package, the methods package). This book does not teach S4 in depth. If you need it, you will know: Bioconductor work, formal class hierarchies, or extending packages that already use S4. For most R programming, S3 or S7 is the right choice.
24.7 S7
S7 is available on CRAN. In May 2024, R-Core merged four patches into base R (version 4.4) that generalize the internal dispatch machinery to support systems like S7. Those patches did not add S7 to base R, but they changed the foundation underneath it: UseMethod() and the method lookup tables now accommodate external OOP systems that follow S7’s functional dispatch model. The long-term plan, stated by the R Consortium OOP Working Group, is to incorporate S7 into base R itself.
S3 attached a string to an object and hoped someone would write the right method. S4 replaced hope with a registry, typed slots, and setClass() declarations. new_class() returns a constructor function. new_generic() returns a function. You register methods by assigning to method(). The entire OOP system is made of functions you call, compose, and pass around — the same primitive this book has been building on since Chapter 5. The objects have typed properties and built-in validation, but the machinery that defines classes, creates generics, and dispatches methods is itself just functions.
Classes are created with new_class():
library(S7)
penguin <- new_class("penguin",
properties = list(
name = class_character,
mass = class_double
),
validator = function(self) {
if (self@mass <= 0) "mass must be positive"
}
)
# penguin is now a constructor function:
p <- penguin(name = "Adelie", mass = 3750)
p
#> <penguin>
#> @ name: chr "Adelie"
#> @ mass: num 3750Properties are typed and validated. You access them with @, like S4 slots, but the syntax is cleaner. The validator returns an error message string (or NULL if valid), rather than calling stop().
Generics and methods use new_generic() and method():
greet <- new_generic("greet", "x")
method(greet, penguin) <- function(x, ...) {
paste("Hello,", x@name)
}
p <- penguin(name = "Adelie", mass = 3750)
p@name
#> [1] "Adelie"
greet(p)
#> [1] "Hello, Adelie"Notice what you get for free: the class constructor (penguin(...)) is generated automatically with named arguments matching the properties. The validator runs on construction. Properties are type-checked. Compare this to the three-function pattern you had to build by hand in Section 24.3.
S7 over S3: properties have declared types (class_character, class_double, class_integer). Validation is built in, not bolted on. The syntax for generics and methods is explicit and clean. And S7 is fully compatible with S3: S7 methods can dispatch on S3 classes, and S3 methods can dispatch on S7 classes.
S7 over S4: simpler syntax. No setClass/setGeneric/setMethod ceremony. Builds on S3 dispatch, which everything already uses. Designed for the modern R ecosystem rather than being a parallel system.
Inheritance uses the parent argument:
diving_penguin <- new_class("diving_penguin",
parent = penguin,
properties = list(
dive_depth = class_double
)
)
emperor <- diving_penguin(name = "Emperor", mass = 23000, dive_depth = 500)
emperor
#> <diving_penguin>
#> @ name : chr "Emperor"
#> @ mass : num 23000
#> @ dive_depth: num 500
greet(emperor)
#> [1] "Hello, Emperor"
# inherited from the penguin methodThe child class inherits the parent’s properties and methods. You can override methods for the child class the same way: method(greet, diving_penguin) <- function(x, ...) ....
For new code, use S7 if you want validated properties and formal structure. Use S3 if you want simplicity and zero dependencies. Both are good choices. S4 is for existing codebases that already use it. Do not start new projects with S4 unless you are writing Bioconductor packages.
Exercises
- Rewrite the
"book"class from the earlier exercise as an S7 class with propertiestitle(character),pages(integer), andyear(integer). Add a validator that checkspages > 0andyear > 0. (You will needlibrary(S7)installed.) - Create an S7 generic
describeand a method for yourbookclass that returns"Title (Year, N pages)". - Create a child class
"ebook"with an additionalformatproperty (character, e.g."epub"or"pdf"). Verify that thedescribemethod is inherited.