library(palmerpenguins)
#>
#> Attaching package: 'palmerpenguins'
#> The following objects are masked from 'package:datasets':
#>
#> penguins, penguins_raw
library(stringr)
library(forcats)
library(lubridate)
#>
#> Attaching package: 'lubridate'
#> The following objects are masked from 'package:base':
#>
#> date, intersect, setdiff, union12 Strings, factors, and dates
You have a column of dates in three different formats, a species column that plots in the wrong order, and a free-text field full of trailing whitespace and inconsistent capitalization. Numbers behave; text, categories, and time do not. This chapter covers the three types that derail an afternoon: strings, factors, and dates. Each has a dedicated tidyverse package (stringr, forcats, lubridate), each comes with traps that catch beginners, and none of them will feel fully tamed by the end of a single chapter. The three topics are independent; skip to whichever one is causing you trouble right now.
We will use the palmerpenguins dataset throughout. Load it now:
12.1 Character vectors
A string in R is a character vector of length 1. There is no special string type: "hello" is character(1), the same kind of atomic vector you met in Section 4.2.
x <- "hello"
typeof(x)
#> [1] "character"
length(x)
#> [1] 1Double quotes and single quotes both work. Pick one and stick with it.
Use double quotes. R’s own style guide does, and so does most code you will encounter in the wild. Single quotes earn their keep inside double-quoted strings: "it's easy".
R uses UTF-8 as its modern string encoding, a scheme that Ken Thompson and Rob Pike designed on a placemat in a New Jersey diner in September 1992, one that is backward-compatible with ASCII, self-synchronizing, and variable-width (one to four bytes per character). If accented characters come out garbled, the file is probably Latin-1; stringr::str_conv() or readr::locale(encoding = "latin1") handles that.
Special characters use backslash escapes: \n (newline), \t (tab), \\ (literal backslash). Compare print() and cat():
print("line one\nline two")
#> [1] "line one\nline two"
cat("line one\nline two")
#> line one
#> line twoprint() shows the escape sequence as literal text; cat() renders it. A small difference, until you spend ten minutes wondering why your output has \n in it instead of a line break.
Base R provides a handful of string tools:
nchar("penguin")
#> [1] 7
paste("Gentoo", "penguin")
#> [1] "Gentoo penguin"
paste0("Gentoo", "penguin")
#> [1] "Gentoopenguin"
sprintf("The %s weighs %d grams", "Gentoo", 5200)
#> [1] "The Gentoo weighs 5200 grams"nchar() counts characters, paste() joins with a space, paste0() joins without, and sprintf() does formatted substitution borrowing syntax from C. These work, but they are inconsistent in argument order and NA handling, and inconsistency compounds fast once you start chaining operations. What would a consistent interface look like?
Exercises
- What does
nchar(NA)return? What aboutnchar("")? - Use
paste()to combine"Species",":", and"Adelie"into a single string. Then do the same withpaste0(). What is different? - Use
sprintf()to produce the string"Island: Biscoe, n = 168".
12.2 stringr: consistent string operations
Base R has grep(), grepl(), sub(), gsub() — they work, but the argument orders are inconsistent, the return types vary, and good luck guessing what gsub stands for. No convention ties them together, so you memorize each one separately.
stringr fixes this with a single rule: every function starts with str_, takes the string as the first argument, and takes the pattern second. Once you know the convention, you can guess function names before you look them up.
The essentials:
str_length("penguin")
#> [1] 7
str_sub("penguin", 1, 4)
#> [1] "peng"
str_c("Gentoo", "penguin", sep = " ")
#> [1] "Gentoo penguin"str_length() counts characters (like nchar()), str_sub() extracts by position, and str_c() combines strings. But unlike paste(), str_c() propagates NA, which means a missing value stays missing instead of silently becoming the literal text "NA":
paste("hello", NA)
#> [1] "hello NA"
str_c("hello", NA)
#> [1] NACase conversion and whitespace cleaning:
str_to_upper("gentoo")
#> [1] "GENTOO"
str_to_title("gentoo penguin")
#> [1] "Gentoo Penguin"
str_trim(" messy data ")
#> [1] "messy data"
str_squish(" too many spaces ")
#> [1] "too many spaces"Pattern matching is where stringr earns its place. Where base R gives you grepl(), stringr gives you str_detect(), a name that actually describes what it does:
species <- penguins$species
str_detect(species, "Gentoo")[1:10]
#> [1] FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSEExtraction and replacement:
islands <- c("Biscoe Island", "Dream Island", "Torgersen Island")
str_extract(islands, "^[A-Za-z]+")
#> [1] "Biscoe" "Dream" "Torgersen"
str_replace(islands, "Island", "Isl.")
#> [1] "Biscoe Isl." "Dream Isl." "Torgersen Isl."Splitting:
str_split("one-two-three", "-")
#> [[1]]
#> [1] "one" "two" "three"str_split() returns a list because each input string could split into a different number of pieces, the same reason strsplit() does in base R, and a preview of why lists matter for irregular output.
On the penguins data:
str_to_upper(levels(penguins$island))
#> [1] "BISCOE" "DREAM" "TORGERSEN"
sum(str_detect(penguins$species, "Adelie"))
#> [1] 152Exercises
- How many penguins have species names containing the letter “e”? Use
str_detect(). - Use
str_sub()to extract the first three letters of each island name inpenguins$island. - Use
str_replace_all()to replace all spaces in"Gentoo penguin species"with underscores.
12.3 Regex essentials
You have seen str_detect() matching literal text. But what if you need to match “any string that starts with a capital letter and ends with a digit”? Literal matching cannot express that. Regular expressions can.
Regular expressions are a mini-language for describing patterns in text. The syntax is not R-specific — the same patterns work in Python, JavaScript, and grep on the command line. What follows covers enough to read and write simple patterns; ?regex has the full reference.
The core building blocks:
| Pattern | Matches |
|---|---|
. |
Any single character |
^ |
Start of string |
$ |
End of string |
[abc] |
Any of a, b, or c |
[0-9] |
Any digit |
+ |
One or more of the preceding |
* |
Zero or more of the preceding |
? |
Zero or one of the preceding |
(A|B) |
A or B (alternation) |
\\d |
Digit (same as [0-9]) |
\\s |
Whitespace |
\\w |
Word character (letter, digit, underscore) |
The doubled backslashes (\\d instead of \d) exist because R strings process backslashes first: to get a literal \d to the regex engine, you write "\\d" in R. Regular expressions themselves come from formal language theory, where Stephen Kleene defined regular languages in 1956 using concatenation, alternation, and closure (repetition). Ken Thompson implemented them in the QED editor in 1968 and grep in 1973; the * quantifier is still called the Kleene star after him.
str_view() lets you see what a pattern matches, making it the best tool for building and debugging:
fruits <- c("apple", "banana", "cherry", "date", "elderberry")
str_view(fruits, "[aeiou]")
#> [1] │ <a>ppl<e>
#> [2] │ b<a>n<a>n<a>
#> [3] │ ch<e>rry
#> [4] │ d<a>t<e>
#> [5] │ <e>ld<e>rb<e>rrySome practical patterns:
# Strings that start with a capital letter
str_detect(c("Hello", "world", "R"), "^[A-Z]")
#> [1] TRUE FALSE TRUE
# Strings that end in a digit
str_detect(c("room101", "lobby", "floor3"), "\\d$")
#> [1] TRUE FALSE TRUE
# Extract numbers from text
str_extract("penguin weighs 5200 grams", "\\d+")
#> [1] "5200"You do not need to memorize regex. Know that it exists, know the basics from the table above, and keep the stringr cheatsheet within arm’s reach. That is enough.
Exercises
- Write a regex that matches strings starting with “G” and ending with “o”. Test it on
c("Gentoo", "Galileo", "Go", "Gusto", "Goo"). - Use
str_extract_all()to pull all words (sequences of\\w+) from"The quick brown fox". - Use
str_detect()and a regex to find which island names inpenguins$islandcontain two consecutive vowels.
12.4 Why factors exist
Strings handle free text. But many columns are not free text: species, island, treatment group. These columns have a fixed set of values, and the order of those values matters for plots and models. That is where factors come in.
Try plotting penguin species as a bar chart. The bars come out in alphabetical order: Adelie, Chinstrap, Gentoo. You want them ordered by count, or by body mass, or by some logic that makes sense for your story. Character vectors do not carry ordering information, so you cannot tell the plot “put Gentoo first” without something more.
A factor is that something more: a vector of integers with labels. When R stores c("male", "female", "female") as a factor, it is really storing c(1, 2, 2) with a mapping (1 = “female”, 2 = “male”, alphabetical by default).
x <- factor(c("male", "female", "female"))
x
#> [1] male female female
#> Levels: female male
typeof(x)
#> [1] "integer"
unclass(x)
#> [1] 2 1 1
#> attr(,"levels")
#> [1] "female" "male"typeof() returns "integer". unclass() strips the factor shell and shows the integers underneath.
In type theory, a factor is a sum type: factor(c("male", "female")) defines a type with exactly two variants, male | female, and a value must be one of them. Logical vectors are the simplest case: Bool = TRUE | FALSE. Factors generalize it: Species = Adelie | Chinstrap | Gentoo is a sum type with three variants. Where a data frame is a product type (combine fields with AND, Chapter 11), a factor is a sum type (choose one variant with OR). The distinction matters because product types grow by adding fields, while sum types grow by adding variants, and the two compose differently. Rust’s enum, Haskell’s algebraic data types, and TypeScript’s union types all make this distinction explicit; R keeps it implicit in the factor machinery, but the structure is the same.
That implicit structure does real work. When you store 10,000 penguin observations with three species, a character vector holds the full string "Adelie" repeated thousands of times. A factor holds the integer 1 and a three-entry lookup table. The compression matters for memory, but it matters more for reasoning: the moment you write factor(species, levels = c("Adelie", "Chinstrap", "Gentoo")), you are declaring a contract. These three values can appear in this column, and nothing else. That contract is what lets lm() generate dummy variables automatically — the full set of categories is declared up front, not discovered by scanning the data. It also means fct_reorder() can reorder levels without inventing ones that do not exist, and data-entry errors ("Adelie" vs "adelie") surface at the moment of assignment rather than downstream in a model that silently treats them as two distinct categories. A character vector is an open set: anything goes. A factor is a closed set, and closed sets are easier to reason about, for you and for every function that receives them.
So why does R have this machinery at all? Statistical models need to encode categorical variables, and factors tell R “these are categories, not arbitrary text.” Without factors, R would not know that "male" and "female" represent a finite set of categories rather than two arbitrary strings.
Historical baggage worth knowing: data.frame() used to convert all strings to factors by default, which caused years of confusion and the ubiquitous stringsAsFactors = FALSE incantation. R 4.0 (released 2020) changed the default to FALSE, ending decades of pain. If you see old code with stringsAsFactors, now you know why it is there.
When do factors actually help?
- Controlling the order of levels in plots (bars, legends, facets)
- Statistical modeling (
lm(),glm(), and friends) - Any time the set of possible values matters: months, Likert scales, treatment groups
When they do not: most data wrangling. If you are filtering and counting, character vectors are fine. But the moment you need a plot where the ordering tells a story, you will reach for factors, and at that point the question becomes how to wrangle them without pain.
Exercises
- Create a factor from
c("low", "medium", "high", "low", "high"). What are the levels? In what order? - Use
unclass()to see the integer codes. Which integer corresponds to “low”? - What happens if you try to assign a value that is not in the levels? Try
x[1] <- "extreme"on your factor.
12.5 forcats: taming factors
Base R’s factor() lets you set levels manually:
sizes <- factor(c("small", "medium", "large"), levels = c("small", "medium", "large"))
sizes
#> [1] small medium large
#> Levels: small medium largeThe levels argument controls the allowed values and their order. Without it, R defaults to alphabetical, which is why “high” comes before “low” and plots look wrong. But manually specifying levels for every factor in every analysis gets tedious fast, and tedium breeds errors.
forcats provides cleaner tools, all starting with fct_:
# Reorder levels manually
fct_relevel(sizes, "large", "medium", "small")
#> [1] small medium large
#> Levels: large medium small# Order levels by frequency in the data
fct_infreq(penguins$species) |> table()
#>
#> Adelie Gentoo Chinstrap
#> 152 124 68# Collapse rare levels into "Other"
fct_lump_n(penguins$species, n = 2) |> table()
#>
#> Adelie Gentoo Other
#> 152 124 68# Rename levels
fct_recode(penguins$species, AP = "Adelie", GP = "Gentoo", CP = "Chinstrap") |> head()
#> [1] AP AP AP AP AP AP
#> Levels: AP CP GPThe most useful function is fct_reorder(), which reorders levels by a summary of another variable, and this is where factors stop being an annoyance and start being a plotting superpower:
library(ggplot2)
penguins_clean <- penguins[!is.na(penguins$body_mass_g), ]
ggplot(penguins_clean, aes(x = fct_reorder(species, body_mass_g, median), y = body_mass_g)) +
geom_boxplot() +
labs(x = "Species (ordered by median body mass)", y = "Body mass (g)")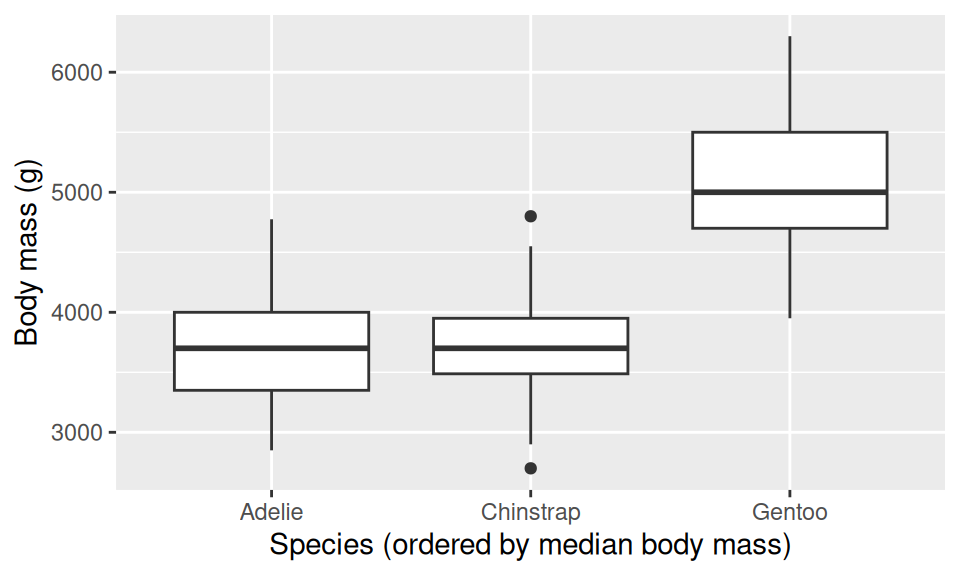
Without fct_reorder(), species appear alphabetically: Adelie, Chinstrap, Gentoo. With it, they appear in order of median body mass, and suddenly the plot tells a story instead of reciting the alphabet. This is the moment factors click: the labels reflect the data, not a sorting convention that has nothing to do with penguins.
Exercises
- Use
fct_infreq()onpenguins$islandto see which island has the most observations. - Use
fct_lump_n()withn = 1onpenguins$species. What happens? - Create a bar chart of
penguins$specieswith bars ordered by frequency (hint:fct_infreq()insideaes()).
12.6 Dates and times
Strings are irregular text; factors are constrained categories. Dates look like strings but behave like numbers, and that dual nature is the source of most of the trouble.
Add one month to January 31. What do you get? February 28? February 31 (which does not exist)? An error? The answer depends on your tool, and the fact that reasonable tools disagree tells you something about why date arithmetic is harder than it looks. Time zones, leap years, daylight saving transitions, and varying month lengths conspire to make even simple calculations surprisingly tricky.
R has three date-time classes:
Date: date only, stored as days since 1970-01-01POSIXct: date + time, stored as seconds since 1970-01-01 (compact, use in data frames)POSIXlt: date + time as a named list of components (rarely needed)
today <- Sys.Date()
today
#> [1] "2026-04-07"
typeof(today)
#> [1] "double"
unclass(today)
#> [1] 20550The number from unclass() is days since the Unix epoch. Dates in R count from 1970-01-01 because Unix measured time as seconds from that date; 32-bit signed integers can count seconds from 1970 until January 19, 2038 (the “Year 2038 problem”), and R, Python, JavaScript, and most databases all inherit this same epoch. Date arithmetic works as you would expect:
as.Date("2026-03-07") - as.Date("2026-01-01")
#> Time difference of 65 daysR returns a difftime object. Subtraction works, and so does addition with scalars:
as.Date("2026-01-01") + 30
#> [1] "2026-01-31"The base R parsing function is as.Date(), which expects ISO 8601 format ("YYYY-MM-DD") by default:
as.Date("2026-03-07")
#> [1] "2026-03-07"
as.Date("07/03/2026", format = "%d/%m/%Y")
#> [1] "2026-03-07"The format argument uses %Y (4-digit year), %m (month), %d (day), and similar codes. These are hard to remember and easy to get wrong, especially when your data mixes formats. Is "03/07/2026" March 7th or July 3rd? The format string decides, and if you pick wrong, R will not complain.
Exercises
- What day number (since 1970-01-01) is today? Use
unclass(Sys.Date()). - What date is 1000 days from today? Use
Sys.Date() + 1000. - How many days are between
"2024-02-28"and"2024-03-01"? (2024 is a leap year.)
12.7 lubridate: dates for humans
Those %Y/%m/%d format codes? lubridate lets you forget them. Its parsing functions are named after the order of components, so the function name is the format:
ymd("2026-03-07")
#> [1] "2026-03-07"
dmy("07/03/2026")
#> [1] "2026-03-07"
mdy("03-07-2026")
#> [1] "2026-03-07"All three produce the same date. No format strings to memorize, no ambiguity about which code means what.
For date-times:
ymd_hms("2026-03-07 14:30:00")
#> [1] "2026-03-07 14:30:00 UTC"Extracting components:
d <- ymd("2026-03-07")
year(d)
#> [1] 2026
month(d)
#> [1] 3
day(d)
#> [1] 7
wday(d, label = TRUE)
#> [1] Sat
#> Levels: Sun < Mon < Tue < Wed < Thu < Fri < SatDate arithmetic with human-readable units:
d + days(30)
#> [1] "2026-04-06"
d + months(1)
#> [1] "2026-04-07"
d + years(1)
#> [1] "2027-03-07"lubridate handles month-length differences correctly: adding one month to January 31 gives February 28 (or 29 in a leap year), not an error. That question from the previous section now has a clear answer.
lubridate distinguishes three kinds of time spans:
- Duration: exact number of seconds.
ddays(1)is always 86400 seconds. - Period: human units.
days(1)is “one day,” which could be 23 or 25 hours around daylight saving transitions. - Interval: an anchored span with a start and end.
For most work, periods (days(), months(), years()) are what you want. Durations (ddays(), dmonths()) matter when you need physical time, the kind a stopwatch measures rather than a calendar.
# How old is R? (first public release: 1993-08-01)
interval(ymd("1993-08-01"), Sys.Date()) %/% years(1)
#> [1] 32Parse with lubridate, extract with lubridate, do arithmetic with lubridate. Reach for base R date functions only when you need zero dependencies.
Exercises
- Parse the following dates:
"15-Jan-2024","2024/06/30","December 25, 2023". Which lubridate function does each need? - What day of the week were you born? Use
ymd()andwday(label = TRUE). - Compute the number of days between
"2020-03-01"and"2026-03-01". Then compute the number of months usinginterval()and%/%.
12.8 Summary
Each of these data types has a matching tidyverse package:
| Data type | Base R | Tidyverse package | Prefix |
|---|---|---|---|
| Text | paste(), grep(), sub() |
stringr | str_ |
| Categories | factor(), levels() |
forcats | fct_ |
| Dates | as.Date(), Sys.Date() |
lubridate | ymd(), year(), … |
The tidyverse packages give these operations a consistent interface and naming scheme. In some cases (like str_detect vs grepl) the difference is mostly cosmetic; in others (like lubridate’s timezone handling) the package catches real bugs that base R silently lets through.
Chapter 11 covered the structure; this chapter covered the three column types that cause the most trouble. But data frames need data, so Chapter 13 deals with getting files off disk and into R. After that, Chapter 14 shows how these pieces combine: str_detect() inside filter(), fct_reorder() inside ggplot(), date arithmetic inside mutate(). That is where the column-level work starts saving you real time.